Abstract
The Eucalyptus loxophleba complex comprises two tree taxa (E. loxophleba ssp. loxophleba and ssp. supralaevis), two mallee taxa (E. loxophleba ssp. lissophloia and ssp. gratiae), and three rare mallee species, of which one, E. blaxellii, is included in this study. The genetic diversity and phylogenetic relationships between the taxa, particularly in relation to habit, were assessed using anonymous nuclear RFLP loci. The level of diversity in the taxa was high and similar to that detected in other eucalypt species. The populations showed low differentiation at both the subspecies and the species levels. Phylogenetic relationships showed some genetic separation between the tree and mallee habit but no separation of the two taxa within the tree habit or within the mallee habit. The genetic analysis does not support the recognition of E. gratiae as a separate species. The geographically restricted E. blaxellii showed similar levels of diversity to populations of the other widespread taxa of E. loxophleba.
Similar content being viewed by others
Introduction
Many woody plants show variation in growth form, often having both tree and woody clump forms. The genus Eucalyptus has tree forms and multistemmed clumps or ‘mallees’, where several sparingly branched stems arise from a main underground root stock (lignotuber) (Wood, 1929). Eucalypts that do not form a lignotuber have a tree growth form, but those that do form a lignotuber can have a mallee habit or a tree habit (Dean, pers. comm.). The presence or absence of a lignotuber appears to be a heritable character and therefore genetically determined (Pryor, 1957; Dean, unpubl.). The habit (tree vs. mallee) of eucalypt species is generally consistent within taxa, although habit can be modified in response to disturbance, particularly fire (Lacey, 1983; Lacey & Johnston, 1990). The mallee habit occurs throughout the genus and is hypothesized to have evolved numerous times (Hill, 1989). Mallees occur in many ecological habitats but are dominant in the semiarid regions and may be an adaptation to stress resulting from climatic conditions (Martin, 1989) or nutrient deficient soils (Main, 1996).
Habit has been used extensively in the classification and taxonomy of eucalypt species and there are species groups where habit is one of the main characters used to define the taxa (e.g. Brooker & Hopper, 1991). However, habit represents a single morphological character and changes in expression of morphological traits can result from small changes in one or a few genes (Gottlieb, 1984; Doebley, 1993). Whilst there have been numerous studies on the morphological and anatomical differences between habits (Mullette, 1978; Bamber & Mullette, 1978; Lacey, 1983; Lacey & Johnston, 1990), the extent of genetic differentiation between related eucalypt taxa with different habits has not been investigated. Genetic differentiation has been observed between growth forms of Picea englemannii and Abies lasiocarpa at sites along the tree line at the edge of the species' distribution in western North America (Grant & Mitton, 1977; Shea & Grant, 1985).
Eucalyptus loxophleba Benth. is a species complex in which all taxa form a lignotuber but where both the mallee and tree habit are present in separate taxa. It is a widespread species distributed across south-west Western Australia, except for the extreme south-west coastal areas (Fig. 1). Eucalyptus loxophleba Benth. has four subspecies: ssp. loxophleba, a tree with rough bark to the upper branches; ssp. supralaevis L.A.S. Johnson & K.D. Hill, a tree with a stocking of rough bark on the trunk and smooth branches; ssp. lissophloia L.A.S. Johnson & K.D. Hill, a smooth-barked mallee; and spp. gratiae Brooker, a smooth-barked mallee with larger buds and fruits than ssp. lissophloia (Hill & Johnson, 1992). When describing ssp. lissophloia and ssp. supralaevis, Johnson & Hill (1992) also raised ssp. gratiae to specific rank, although this revision is not generally accepted and will not be followed here. The mallee subspecies are found in the south and east of the distribution in south-west Western Australia, and the trees in the west and north of the distribution (Fig. 1). In addition to differentiation in habit, the subspecies also show differences in leaf chemistry as the two mallee taxa both possess high levels of the leaf oils, 1,8-cineole and 4-methylpent-2-yl acetate (Grayling, 1996).
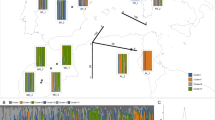
Distributions of Eucalyptus loxophleba and E. blaxellii and the location of sampled populations. Population numbers according to Table 1.
There are three other mallee species that are related to E. loxophleba, and all have highly restricted distributions. The most closely related species based on morphological similarity, E. blaxellii Johnson & Hill, occurs in the Moresby Range north of Geraldton, within the distribution of ssp. loxophleba and ssp. supralaevis. The other two species, E. semota Macph. & Grayling and E. articulata Brooker & Hopper, occur in the inland desert region outside the range of E. loxophleba.
In this study, RFLP analysis of the nuclear genome was used to investigate the genetic relationships within the E. loxophleba complex, particularly the level of differentiation between the tree and mallee taxa.
Materials and methods
Plant material
Leaf samples were collected from 10 individuals, from each of 20 populations. Populations where selected to represent the geographical distribution of each taxon (Fig. 1). Two populations were also selected as ‘intermediate’ populations, bearing features of more than one taxon (Hill & Johnson, 1992; Grayling, 1996). A population of the related mallee species, E. blaxellii was also collected. Five individuals from the Wagin population of E. spathulata, which is in a different series to E. loxophleba, were included as an outgroup in the study.
DNA was extracted from the leaves of the 205 individuals following the methods outlined in Byrne et al. (1998). DNA samples were digested with BglII, DraI, or EcoRV, Southern blotted and hybridized with 30 nuclear RFLP probes from the eucalypt genetic linkage map (Byrne et al., 1995; c030, c092, c113, c115, c116, c135, c170, c238, c299, c333, c395, c411, c451, g059, g067, g086, g095, g099, g142, g154, g174, g183, g195, g233, g243, g250, g256, g261, g425, g474). Hybridization was according to Byrne & Moran (1994) and the probes were hybridized against DNA digested with one of the three enzymes used. Any fragment pattern not able to be scored using one enzyme–probe combination was subsequently re-assayed with another of the enzymes.
Data analysis
Alleles were numbered at each locus according to the size of the fragment, with the largest fragment designated allele 1. Allelic diversity parameters were calculated using POPGENE (Yeh et al., 1997). UPGMA analysis was carried out, based on unbiased genetic distance measures (Nei, 1978). The strength of the population clusters within the UPGMA was examined using three-dimensional SSH (semistrong multi-dimensional scaling) ordination, based on the same genetic distance measures, using the computer program PATN (Belbin, 1993). A phylogenetic analysis of population relationships was carried out using a continuous character maximum likelihood algorithm in PHYLIP (Felsenstein, 1993) using E. spathulata as the outgroup. Bootstrap support for nodes in the UPGMA and maximum likelihood analyses was estimated from 100 replications using the Seqboot algorithm in PHYLIP (Felsenstein, 1993).
Results
Information derived from 25 probes was used in the analysis, because five probes produced banding patterns that could not be reliably scored. Polymorphism in E. loxophleba was high, with only two loci, c115 and g195, monomorphic in all populations. Fourteen loci were polymorphic in all populations, and nine loci were monomorphic in one or more populations. The number of alleles detected at a locus ranged from one to 21 with a maximum number of 13 alleles in any one population. The frequency of alleles was skewed slightly towards alleles of low frequency; 33% of alleles had frequencies of < 0.1, and 22% of alleles had frequencies > 0.5.
The allelic diversity measures for each population of E. loxophleba, and means over all populations and for each taxon, are presented in Table 1. The values for mean number of alleles per locus (A), mean number of polymorphic loci (P), observed heterozygosity (Ho), and gene diversity (He), were all high and similar both between populations and between taxa, although ssp. loxophleba had consistently lower values than the other taxa.
The fixation index, FIS, was small and positive for most populations but not significantly different from zero indicating minimal inbreeding. At the taxon level FIS was higher in ssp. loxophleba but again none of the values were significantly different from zero. Differentiation between populations (FST) was low, at both the subspecies (0.046–0.081) and species (0.089) level, with no strong differentiation evident within the species. Differentiation was lowest in ssp. gratiae, reflecting the smaller geographical distribution of this taxon.
The population of E. blaxellii showed similar levels of genetic diversity to populations of E. loxophleba (Table 1). The population of E. spathulata had lower levels of diversity than populations of E. loxophleba, due to the smaller sample size for this population (five individuals vs. 10). The fixation index, FIS, was close to zero for both species.
Unbiased estimates of genetic distance between all pairwise comparisons of populations of E. loxophleba were low, with the highest distance (0.105) occurring between the ssp. lissophloia population at The Pimple and the ssp. loxophleba population at Geraldton. The lowest genetic distance (0.014) occurred between the ssp. lissophloia population at The Pimple and ssp. gratiae at Dumbleyung. The average genetic distance between E. blaxellii and populations of E. loxophleba (0.089) was low for separation between species as distances between conspecific populations are generally below 0.1 (Crawford, 1989). For E. spathulata, the genetic distance from populations of E. loxophleba was large (0.222) and the distance to E. blaxellii larger still (0.247).
UPGMA analysis based on genetic distance shows the main group of E. loxophleba populations split into two clusters, although the genetic distance separating the clusters is small (D=0.057; Fig. 2). One cluster contains populations of the mallee taxa, ssp. lissophloia and ssp. gratiae, and the other contains populations of the tree taxa, ssp. loxophleba and ssp. supralaevis, with neither group showing separation of the taxa within them. The two pairs of tree and mallee populations that are geographically closest (ssp. lissophloia at Narembeen and ssp. supralaevis at Koorda; ssp. gratiae at Dumbleyung and ssp. loxophleba at Kulin) do not show close genetic distance. In each case the tree or mallee population is more similar to other populations of the same habit than to the geographically closest population of the alternate habit. There are two populations placed outside the two main clusters of populations. They are geographically distant populations of the tree taxa occurring in the northernmost and southernmost parts of the range (ssp. loxophleba at the Stirling Range and ssp. supralaevis at Murchison River). The two populations do not show strong genetic similarity (D=0.063) and both are distant from the remainder of the populations (D=0.069). The level of bootstrap support for clusters within E. loxophleba was low (4–39%), except for three pairs of populations with 60–82% bootstrap support. Eucalyptus blaxellii is separate from the populations of E. loxophleba (with strong bootstrap support of 83%) and E. spathulata is quite distinct (bootstrap support 100%).
The SSH ordination plotted in three dimensions produced a stress value of 0.125 after 22 iterations, indicating a fair fit between the ordination and the data. The ordination in two dimensions is presented in Fig. 3. The relationship between populations in the ordination plot shows a broad spread of the E. loxophleba populations in both dimensions. The mallee populations are grouped together with some separation from the tree populations on the x- and y-axes. The Stirling Range and Murchison River populations show little affinity to one another, and are distinct from the rest of the ssp. loxophleba and supralaevis populations in both dimensions and separate from the mallee populations on the y-axis. There was prominent separation of E. blaxellii on both the x-and y-axis.
Ordination analysis of genetic distance between populations of Eucalyptus loxophleba and E. blaxellii. Populations are represented by numbers (see Table 1). The circles surrounding populations indicate the tree (dotted line) and mallee (dashed line) populations of E. loxophleba.
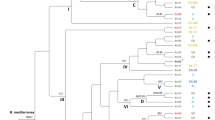
A continuous character maximum likelihood analysis of population phylogeny separated ssp. loxophleba and ssp. supralaevis populations from ssp. lissophloia and ssp. gratiae populations (Fig. 4). Eucalyptus blaxellii was not separate from E. loxophleba and was placed in the cluster of tree populations. The intermediate population at Harrismith fell between the cluster of tree populations and the mallee populations. The populations of tree forms were grouped into a specific cluster compared to the mallee populations in which the cluster was less well defined. The bootstrap support for all nodes was low (6–63%) and indicated no support for the differentiation of the mallee and tree clusters.
Discussion
The E. loxophleba complex is a highly variable group that showed no clear genetic differentiation between the described taxa although there was limited evidence for genetic separation of the mallee and tree habit. The level of allelic diversity present in the E. loxophleba complex is similar to that found in other eucalypt species that have been studied using similar sampling strategies and the same RFLP probes, E. kochii (Byrne, 1999), E. nitens (Byrne et al., 1998) and E. camaldulensis (Butcher et al., 2001). Although there was high diversity within the nuclear genome of E. loxophleba there was little differentiation between populations. Eucalyptus loxophleba ssp. loxophleba and ssp. supralaevis showed a greater level of differentiation between populations than ssp. lissophloia and ssp. gratiae, which would be expected as they cover a larger geographical range. A high level of diversity with little population differentiation was also detected in E. kochii (Byrne, 1999) and E. camaldulensis (Butcher et al., 2001).
There was no clear differentiation of the taxa in E. loxophleba, with no separation between populations of the mallee taxa, ssp. lissophloia and ssp. gratiae, or between populations of the tree taxa, ssp. loxophleba and ssp. supralaevis. However, there was some evidence of separation of the mallee populations from the tree populations in all the analyses although the patterns were not always consistent, and the level of confidence support was low. The distance and ordination analyses showed a tighter cluster of the mallee form than the tree form with some divergence of populations of the tree form. In contrast, the maximum likelihood analysis showed a tighter cluster of populations of the tree form than those of the mallee form. The level of differentiation between the mallee and tree habits suggests that the evolution of the mallee habit in this species may be recent in evolutionary history, and the influence of common ancestry is still evident in the overall similarity within the species.
The lack of separation between the populations of ssp. gratiae and those of ssp. lissophloia, is consistent with previous analyses of both morphology and leaf oil composition where there was no clear separation between these taxa, but rather a clinal variation across the range (Grayling, 1996). Based on this work, Grayling (1996) proposed the maintenance of two taxa, but with the demotion of ssp. gratiae from specific rank to subspecies level. The genetic data presented here indicate that the mallee forms comprise a single genetic entity and maintenance of two separate mallee taxa may be unwarranted.
Similarly to the mallees, there was a lack of genetic separation between most of the populations of the two tree taxa, ssp. loxophleba and ssp. supralaevis. The two populations that were genetically distinct from the main cluster of tree populations in the distance analysis were the northern-most and southern-most populations sampled. Populations at the northern end of the range of ssp. supralaevis show some morphological variation, having stouter trunks and branch angles nearing horizontal. It is likely that the more distinct morphological and genetic characteristics of the northern population represent divergence at the edge of the species' range.
The second genetically distinct population in the distance analysis was that collected from the Stirling Range. The Stirling Range is of a different geology and higher elevation to that of the surrounding areas, and the area is noted for the high level of local endemism in the flora (Hopkins et al., 1983; Hopper et al., 1996). This area is the southern extent of the range of the E. loxophleba complex and it is likely that divergence at the edge of the range is also the cause of the differentiation of this population from those in the main part of the range. No distinctive differences in morphology or leaf oil composition of another nearby population in the Stirling Range were noted by Grayling (1996). The lack of distinctness of both the Stirling Range and Murchison River populations in the maximum likelihood analysis also suggests that the differences in these populations represent recent rather than ancestral variation.
Two intermediate populations with features of more than one taxon were included in the study. The population at Harrismith has the mallee habit but has a basal stocking of rough bark that is characteristic of ssp. loxophleba. It occurs within the ranges of three taxa, ssp. loxophleba, ssp. lissophloia and ssp. gratiae. This population may be a population of ssp. loxophleba in which disturbance, such as a severe fire, has invoked the mallee habit; however, it did not cluster with the tree forms in either of the phenetic analyses indicating that it does not share genetic similarity with other ssp. loxophleba populations. The position of the population between the mallee and tree clusters in the maximum likelihood analysis and its similarity with the mallee populations in the distance analysis suggests that it represents an intermediate population in which the differentiation of morphological characters is not complete.
The other intermediate population at Wave Rock is morphologically ssp. loxophleba but has an oil composition characteristic of ssp. lissophloia. It is an outlier to the main range of ssp. loxophleba and occurs within the range of ssp. lissophloia. This population clustered with the tree forms in both the phenetic and phylogenetic analyses confirming its morphological identity as a population of ssp. loxophleba.
The low genetic distance between Eucalyptus blaxellii and E. loxophleba is consistent with them being closely related species and suggests that they may represent recent speciation (Crawford, 1989). Eucalyptus blaxellii showed greater similarity to the mallee forms of E. loxophleba than to the tree forms which is consistent with the mallee habit of E. blaxellii, even though it is geographically closer to the tree forms than to the mallee forms. However, in the maximum likelihood analysis, E. blaxellii showed a stronger relationship to the tree forms than the mallee forms, which most likely reflects coancestry with nearby populations of ssp. loxophleba and ssp. supralaevis.
The pattern of genetic diversity in E. loxophleba showed some evidence for genetic differentiation between the tree and mallee habit, but this differentiation was not strong. However, the intermediate populations with a mallee or tree habit but other traits characteristic of taxa of the alternate habit, showed genetic similarity to other populations based on their habit rather than other traits. This indicates that habit is representative of genetic relatedness and that habit is a more reliable character for identification than other traits that characterize the taxa. Furthermore, the closely related mallee species E. blaxellii showed greater genetic similarity to the mallee form than to the tree form of E. loxophleba.
References
Bamber, R. K. and Mullette, K. J. (1978). Studies of the lignotubers of Eucalyptus gummifera (Gaertn & Hochr.). II: Anatomy. Aust J Bot, 26: 15–22.
Belbin, L. (1993). PATN pattern analysis package. CSIRO Publications, Canberra.
Brooker, M. I. H. and Hopper, S. D. (1991). A taxonomic revision of Eucalyptus wandoo, E. redunca and allied species (Eucalyptus series Levispermae Maiden – Myrtaceae) in Western Australia. Nuytsia, 8: 1–189.
Butcher, P. A., Otero, A., Mcdonald, M. W. and Moran, G. F. (2001). Nuclear RFLP variation in Eucalyptus camaldulensis Dehnh. from Northern Australia. Heredity. submitted.
Byrne, M. (1999). High genetic identities between three oil mallee taxa, Eucalyptus kochii ssp. kochii, ssp. plenissima and E. horistes, based on nuclear RFLP analysis. Heredity, 82: 205–211.
Byrne, M. and Moran, G. F. (1994). Population divergence in the chloroplast genome of Eucalyptus nitens. Heredity, 73: 18–28.
Byrne, M., Murrell, J. C., Allen, B. and Moran, G. F. (1995). An integrated genetic linkage map for eucalypts using RFLP, RAPD and isozyme markers. Theor Appl Genet, 91: 869–875.
Byrne, M., Parrish, T. L. and Moran, G. F. (1998). Nuclear RFLP diversity in Eucalyptus nitens. Heredity, 81: 225–232.
Crawford, D. J. (1989). Enzyme electrophoresis and plant systematics. In: Soltis, D. E. and Soltis, P. S. (eds) Isozymes in Plant Biology, pp. 146–164. Dioscorides Press, Portland, OR.
Doebley, J. (1993). Genetics, development and plant evolution. Curr Opin Genet Devel, 3: 865–872.
Felsenstein, J. (1993). PHYLIP (Phylogeny Inference Package), v.3.5c. Department of Genetics, The University of Washington, Seattle, WA.
Gottlieb, L. D. (1984). Genetics and morphological evolution in plants. Am Nat, 123: 681–709.
Grant, M. C. and Mitton, J. B. (1977). Genetic differentiation among growth forms of Englemann spruce and subalpine fir at tree line. Arctic Alpine Res, 9: 259–263.
Grayling, P. M. (1996). Chemical and morphological studies of Eucalyptus species and hybrids containing the non-terpenoid ester 4-methylpent-2-yl acetate. Ph.D. Thesis, University of Western Australia.
Hill, K. D. (1989). Mallee Eucalypt communities: their classification and biogeography. In: Noble, J. C. and Bradstock, R. A. (eds) Mediterranean Landscapes in Australia, Mallee Ecosystems and Their Management, pp. 93–108. CSIRO Publications, Melbourne.
Hill, K. and Johnson, L. (1992). Systematic studies in the eucalypts. 5. New taxa and combinations in Eucalyptus (Myrtaceae) in Western Australia. Telopea, 4: 561–634.
Hopkins, A. J. M., Keighery, G. J. and Marchant, N. G. (1983). Species rich uplands of south Western Australia. Proc Ecol Soc Aust, 12: 15–56.
Hopper, S. D., Harvey, M. S., Chappill, J. A., Main, A. R. and Main, B. Y. (1996). The Western Australian biota as Gondwanan heritage – a review. In: Hopper, S. D., Chappill, J. A., Harvey, M. S. and George, A. S. (eds) Gondwanan Heritage. Past, Present and Future of the Western Australian Biota, pp. 1–46. Surrey Beatty, Chipping Norton.
Lacey, C. J. (1983). Development of large plate-like lignoubers in Eucalyptus botryoides Sm. in relation to environmental factors. Aust J Bot, 31: 105–118.
Lacey, C. J. and Johnston, R. D. (1990). Woody clumps and clumpwoods. Aust J Bot, 38: 299–334.
Main, A. R. (1996). Ghosts of the past: Where does environmental history begin? Envir History, 2: 97–114.
Martin, H. A. (1989). Evolution of mallee and its environment. In: Noble, J. C. and Bradstock, R. A. (eds) Mediterranean Landscapes in Australia, Mallee Ecosystems and Their Management, pp. 83–92. CSIRO Publications, Melbourne.
Mullette, K. J. (1978). Studies of the lignotubers of Eucalyptus gummifera (Gaertn. & Hochr.). I. The nature of the lignotuber. Aust J Bot, 26: 9–13.
Nei, M. (1978). Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics, 89: 583–590.
Pryor, L. D. (1957). The inheritance of some characters in Eucalyptus. Proc Linn Soc NSW, 82: 199–200.
Shea, K. L. and Grant, M. C. (1985). Clonal growth in spire-shaped Englemann spruce and subalpine fir trees. Can J Bot, 64: 255–261.
Wood, J. G. (1929). Floristics and ecology of the mallee. Trans R Soc S Aust, 53: 359–437.
Yeh, F. C., Yang, R. -C., Boyle, B. J., Ye, Z. -H. and Mao, J. X. (1997). POPGENE, the user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Centre, University of Alberta, Canada.
Acknowledgements
We thank the Farm Forestry Program of the Department of Conservation and Land Management for helpful discussions and assistance with leaf collections. We are grateful to CSIRO Forestry and Forest Products for the use of the RFLP probes.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hines, B., Byrne, M. Genetic differentiation between mallee and tree forms in the Eucalyptus loxophleba complex. Heredity 87, 566–572 (2001). https://doi.org/10.1046/j.1365-2540.2001.00950.x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1046/j.1365-2540.2001.00950.x
Keywords
This article is cited by
-
Genomic data inform conservation of rare tree species: clonality, diversity and hybridity in Eucalyptus series in a global biodiversity hotspot
Biodiversity and Conservation (2021)
-
Flowering phenology in a Eucalyptus loxophleba seed orchard, heritability and genetic correlation with biomass production and cineole: breeding strategy implications
Scientific Reports (2020)
-
Association genetics of essential oil traits in Eucalyptus loxophleba: explaining variation in oil yield
Molecular Breeding (2017)
-
Genetic entities and hybridisation within the Acacia microbotrya species complex in Western Australia
Tree Genetics & Genomes (2015)
-
Regional genetic differentiation in Western Australian sandalwood (Santalum spicatum) as revealed by nuclear RFLP analysis
Theoretical and Applied Genetics (2003)