Abstract
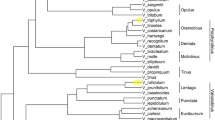
Campomanesia is a well-circumscribed genus within Myrtaceae, but the boundaries among some of its species remain poorly understood. Microsatellite amplification profiles have proven useful DNA fingerprinting for plant species identification, but have never been applied to the taxonomy of this genus. Here we studied 12 species of Campomanesia, including a set of related and non-related species, as well as species with doubtful boundaries. Forty-eight morphological characters and 50 primers were used to test the species. A cluster analysis followed by a group sharpness test and a heat map analysis was performed with both sets of data. A regression tree was constructed using only on the molecular data. Seven groups based on morphological characters and five groups based on molecular characters were revealed by cluster analysis and the sharpness group test. The microsatellite amplification profiles were efficient at distinguishing the studied species, despite the smaller number of distinct groups formed by this marker compared with the morphological data. Among the 50 primers included, 13 were informative for species distinction, and 10 were indicated by the regression tree to species identification. Microsatellite amplification profiles proved to be an informative tool for the taxonomy of Campomanesia.




Similar content being viewed by others
References
Borcard D, Gillet F, Legendre P (2011) Numerical ecology with R. Springer, New York
Chae WB, Hong SJ, Gifford JM, Rayburn AL, Sacks EJ, Juvik JA (2014) Plant morphology, genome size, and SSR markers differentiate five distinct taxonomic groups among accessions in the genus Miscanthus. GCB Bioenergy 6:646–660. https://doi.org/10.1111/gcbb.12101
Christ JA, Hollunder RK, Carvalho MS, Ferreira MF, Garbin ML, Carrijo TT (2018) DNA fingerprinting based on SSR amplification profiles for Piper species identification (Piperaceae). Acta Bot Brasil 32:511–520. https://doi.org/10.1590/0102-33062017abb0268
Crawley MJ (2007) The R Book. Wiley, West Sussex
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Duminil J, Di Michele M (2009) Plant species delimitation: a comparison of morphological and molecular markers. Plant Biosyst 143:528–542. https://doi.org/10.1080/11263500902722964
Galili T, Mitelpunkt A, Shachar N, Marcus-Kalish M, Benjamini Y (2014) Categorize, cluster and classify: a 3-C strategy for scientific discovery in the Medical Informatics platform of the Human Brain Project. In: Discovery science, pp 73–86
Landrum LR (1986) Campomanesia, Pimenta, Blepharocalyx, Legrandia, Acca, Myrrhinium and Luma (Myrtaceae). Flora Neotropica Monograph 45:7–72
Landrum LR (2001) Two new species of Campomanesia (Myrtaceae) from Espírito Santo and Bahia, Brazil. Brittonia 53:534–538
Legendre P, Legendre L (2012) Numerical ecology, 3rd edn. Elsevier, Amsterdam
Lima DF, Mauad AVS, Silva-Pereira V, Smidt EC, Goldenberg R (2015) Species boundaries inferred from ISSR markers in the Myrcia laruotteana complex (Myrtaceae). Plant Syst Evol 301:353–363. https://doi.org/10.1007/s00606-014-1078-9
Luber J, Oliveira MIU, Ferreira MFS, Carrijo TT (2017) Flora do Espírito Santo: Campomanesia (Myrtaceae). Rodriguésia 68:1767–1790. https://doi.org/10.1590/2175-7860201768514
Maechler M, Rousseeuw P, Struyf A, Hubert M, Hornik K (2016) Cluster: cluster analysis basics and extensions. R package version 2.0.4
Nogueira AM, Ferreira A, da Silva-Ferreira MF (2016) Transferability of Microsatellites from Psidium guajava to Eugenia, Myrciaria, Campomanesia, and Syzygium Species (Myrtaceae). Plant Mol Bio Rep 34:249–256
Oksanen J, Blanchet FG, Kindt R (2016) Vegan: community ecology package. R package version 1.17-1. 2010. http://CRAN.R-project.org/package=vegan. Accessed 20 Oct 2018
Oliveira MIU (2013) Estudos taxonômicos e populacionais em Campomanesia Ruiz & Pavón (Myrtaceae, Myrteae), com ênfase no “Complexo Campomanesia xanthocarpa”. Thesis. Universidade Estadual de Feira de Santana, Bahia
Oliveira MIU, Funch LS, Landrum LR (2012) Flora da Bahia: Campomanesia (Myrtaceae). Sitientibus 12:91–107
Pillar VD (1999) How sharp are classifications? Ecology 80:2508–2516. https://doi.org/10.1890/0012-9658
Pillar VD (2006) MULTIV, multivariate exploratory analysis, randomization testing and bootstrap resampling. User’s Guide v. 2.4. http://ecoqua.ecologia.ufrgs.br/. Accessed 20 Oct 2018
R Core Team (2016) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. Accessed 20 Oct 2018
Ripley B (2016) tree: Classification and Regression Trees. R package version 1.0-37. https://CRAN.R-project.org/package=tree. Accessed 20 Oct 2018
Samah S, De Pardo CVT, Cruz MAS, Valadez-Moctezuma E (2016) Genetic diversity, genotype discrimination, and population structure of Mexican Opuntia sp., determined by SSR markers. Plant Mol Bio Rep 34:146–159. https://doi.org/10.1007/s11105-015-0908-4
Satya P, Paswan PK, Ghosh S, Majumdar S, Ali N (2016) Confamiliar transferability of simple sequence repeat (SSR) markers from cotton (Gossypium hirsutum L.) and jute (Corchorus olitorius L.) to twenty-two Malvaceous species. 3 Biotech 6:65. https://doi.org/10.1007/s13205-016-0392-z
Sharawy S, Karakish E (2015) Taxonomic relationships of some species of Orobanche L. evidence from RAPD-PCR and ISSR markers. Pak J Bot 47:437–452
Spooner DM (2016) Species delimitations in plants: lessons learned from potato taxonomy by a practicing taxonomist. J Syst Evol 54(3):191–203. https://doi.org/10.1111/jse.12203
Tóth G, Gáspari Z, Jurka J (2000) Microsatellites in different eukaryotic genomes: survey and analysis. Genome Res 10:967–981
Tuler AC, Carrijo TT, Nóia LR, Ferreira A, Peixoto AL, da Silva Ferreira MF (2015) SSR markers: a tool for species identification in Psidium (Myrtaceae). Mol Biol Rep 42:1501–1513. https://doi.org/10.1007/s11033-015-3927-1
Weber JL, May PE (1989) Abundant class of human DNA polymorphisms which can be typed using the Polymerase Chain Reaction. Am J Hum Genet 44:388–396
Zhang Y, Yuan X, Teng W, Chen C, Wu J (2016) Identification and Phylogenetic Classification of Pennisetum (Poaceae) Ornamental Grasses Based on SSR Locus Polymorphisms. Plant Mol Biol Rep 34:1181–1192. https://doi.org/10.1007/s11105-016-0990
Acknowledgements
We thank Universidade Federal do Espírito Santo for logistical support; the managers and employees of the Conservation Units for their support during visits; the owners of private areas for permission to collect on their properties; Fundação de Amparo à Pesquisa e Inovação do Espírito Santo - FAPES for the fellowship granted to the first author; and the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for the grants to T.T. Carrijo and M.F.S. Ferreira. This study was financed in part by the Coordernação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001 to the first and second authors.
Author information
Authors and Affiliations
Contributions
JL and TTC designed and conducted the study and wrote the article. JL, JAC, and MFS Ferreira conceived and conducted the molecular marker analysis. All authors equally contributed to the editing and revision of the manuscript and approved the final version for submission.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Luber, J., Christ, J.A., Ferreira, M.F.d. et al. Species delimitation within Campomanesia (Myrtaceae) using morphology and amplification profiles of microsatellite markers. Braz. J. Bot 43, 131–137 (2020). https://doi.org/10.1007/s40415-020-00583-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-020-00583-x