Abstract
Background
Among abiotic stresses, soil salinity is one of the major global constraints to growth and productivity in most of the crop plants, limiting current and future agricultural sustainability. Halophytes are the most suitable host plants to dissect the salinity tolerance phenomenon in natural environment. To elucidate the mechanisms of salinity tolerance is an ever growing research area for genetic engineering of crops to improve the salinity tolerance and ultimately to improve the crop yield and quality.
Methods and results
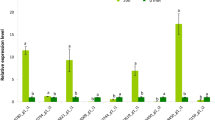
In the present investigation, two un-explored halophytes, i.e. moderately salt tolerant (Dichanthium annulatum) and extremely salt tolerant (Urochondra setulosa) were studied for unrevealing the contributory role of antioxidative system in salinity tolerance at higher saline intensities of ECe ~30, 40 and 50dSm− 1. Experimental findings indicated that the accumulation of hydrogen peroxide (H2O2) content, Superoxide dismutase (SOD) and ascorbate peroxidase (APX) activities were higher in U. setulosa at all saline treatments. Whereas, higher order of malondialdehyde (MDA) content and catalase (CAT) activity was observed in D. annulatum, although the specific enzyme activities of chemically reactive oxygen species (ROS) system increased with increasing levels of salinity in both the halophytes. This differential physiological mechanism was collaborated with the transcriptomic data generated through High throughput sequencing on Illumina platform. Transcriptome expression analysis depicting that a total of 276 and 66 genes coding for various components of ROS system like antioxidant activity, cell redox and glutathione metabolism were differentially expressed in response to salinity stress in U. setulosa and D. annulatum, respectively. Biochemical analysis indicated that H2O2 is detoxified by increased activities of SOD, APX and CAT in D. annulatum, however, in U. setulosa, POX takes over catalase to remove the higher accumulation of H2O2 along with dehydroascorbate reductase (DHAR) and monodehydroascorbate reductase (MDHAR) significantly associated with the differentially expressed transcripts.
Conclusions
The differentially expressed genes for ROS enzymes and antioxidants clearly differentiate and support the detoxification of H2O2 and survival mechanism of Urochondra setulosa and Dichanthium annulatum at different salinity levels. This study provides the reference information for key regulatory genes responsible for salinity stress tolerance and can be used for increasing salinity tolerance of related crop species.








Similar content being viewed by others
Data Availability
The raw data is available with the corresponding author which can be shared on request. Transcriptomics data is submitted at https://www.ncbi.nlm.nih.gov/bioproject/PRJNA561259 and https://www.ncbi.nlm.nih.gov/bioproject/PRJNA665324. The data of differentially expressed genes (DEGs) of both the plants have been deposited in the repository at Mendeley data (https://data.mendeley.com//datasets/c9zwjncxb4/1). Differential antioxidant data has been deposited in the repository at Mendeley data (https://data.mendeley.com/datasets/xd698v9w9s).
References
Abd Elgawad H, Zinta G, Hegab MM et al (2016) High salinity induces different oxidative stress and antioxidant responses in maize seedlings organs. Front Plant Sci 7:276
Aebi H (1984) Catalase in vitro. Methods Enzy. Elsevier, pp 121–126
Ahanger MA, Agarwal RM (2017) Salinity stress induced alterations in antioxidant metabolism and nitrogen assimilation in wheat (Triticum aestivum L) as influenced by potassium supplementation. Plant Physiol Biochem 115:449–460
Ahmad P, Hashem A, Abd-Allah EF et al (2015) Role of Trichoderma harzianum in mitigating NaCl stress in indian mustard (Brassica juncea L) through antioxidative defense system. Front Plant Sci 6:868
Al Hassan M, Chaura J, Donat-Torres MP et al (2017) Antioxidant responses under salinity and drought in three closely related wild monocots with different ecological optima. AoB Plants 9:plx009
Amin I, Rasool S, Mir MA et al (2021) Ion homeostasis for salinity tolerance in plants: a molecular approach. Physiol Plant 171:578–594
Andrews S, Fast QCA (2015) A quality control tool for high throughput sequence data. 2010. https://www.bioinformatics.babraham.ac.uk/
Anschütz U, Becker D, Shabala S (2014) Going beyond nutrition: regulation of potassium homoeostasis as a common denominator of plant adaptive responses to environment. J Plant Physiol 171:670–687
Bankaji I, Cacador I, Sleimi N (2016) Assessing of tolerance to metallic and saline stresses in the halophyte Suaeda fruticosa: the indicator role of antioxidative enzymes. Eco Ind 64:297–308
Beauchamp C, Fridovich I (1971) Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem 44:276–287
Bose JK et al (2014) ROS homeostasis in halophytes in the context of salinity stress tolerance. J Exp Bot 65(5):1241–1257. https://doi.org/10.1093/jxb/ert430
Çelik Ö, Çakır BC, Atak Ç (2019) Identification of the antioxidant defense genes which may provide enhanced salt tolerance in Oryza sativa L. Physiol Mol Biol Plants 25:85–99
Covarrubias AA, Reyes JL (2010) Post-transcriptional gene regulation of salinity and drought responses by plant microRNAs. Plant Cell Environ 33:481–489. https://doi.org/10.1111/j.1365-3040.2009.02048.x
Das AB, Strasser RJ (2013) Salinity-induced genes and molecular basis of salt-tolerant strategies in Mangroves. Molecular stress physiology of plants. Springer, pp 53–86
Dayal D, Mangalassery S, Kumar A (2015) Banni grasslands of Kachchh, Gujarat, India: problems, present status and prospects. In: Ghosh PK, Mahanta SK, Singh JB, Pathak PS (eds) Grassland: a global resource perspective. Range Management Society of India, Jhansi, India
De Carvalho JF, Poulain J, Da Silva C et al (2013) Transcriptome de novo assembly from next-generation sequencing and comparative analyses in the hexaploid salt marsh species Spartina maritima and Spartina alterniflora (Poaceae). Heredity 110:181
Desai AP, Desai S (2019) UV spectroscopic method for determination of vitamin C (ascorbic acid) content in different fruits in south Gujarat region. Int J Environ Sci Nat Res 21
Diray-Arce J, Clement M, Gul B et al (2015) Transcriptome assembly, profiling and differential gene expression analysis of the halophyte Suaeda fruticosa provides insights into salt tolerance. BMC Genomics 16:353
Fan P, Nie L, Jiang P et al (2013) Transcriptome analysis of Salicornia europaea under saline conditions revealed the adaptive primary metabolic pathways as early events to facilitate salt adaptation. PLoS ONE 8:e80595
Fu L, Niu B, Zhu Z et al (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152
Guo S-M, Tan Y, Chu H-J et al (2019) Transcriptome sequencing revealed molecular mechanisms underlying tolerance of Suaeda salsa to saline stress. PLoS ONE 14
Halliwell B, Foyer CH (1978) Properties and physiological function of a glutathione reductase purified from spinach leaves by affinity chromatography. Planta 139:9–17
Heath RL, Packer L (1968) Photoperoxidation in isolated chloroplasts: I. Kinetics and stoichiometry of fatty acid peroxidation. Arch Biochem Biophy 125:189–198
Hossain MA, Asada K (1984) Purification of dehydroascorbate reductase from spinach and its characterization as a thiol enzyme. Plant Cell Physiol 25:85–92
Inan G, Zhang Q, Li P et al (2004) Salt cress. A halophyte and cryophyte Arabidopsis relative model system and its applicability to molecular genetic analyses of growth and development of extremophiles. Plant Physiol 135:1718–1737
Jha B, Agarwal PK, Reddy PS et al (2009) Identification of salt-induced genes from Salicornia brachiata, an extreme halophyte through expressed sequence tags analysis. Genes Genetic Sys 84:111–120
Kohli SK, Khanna K, Bhardwaj R et al (2019) Assessment of subcellular ROS and NO metabolism in higher plants: multifunctional signaling molecules. Antioxidants 8:641
Krasensky J, Jonak C (2012) Drought, salt, and temperature stress induced metabolic rearrangements and regulatory networks. J Exp Bot 63:1593–1608. https://doi.org/10.1093/jxb/err460
Kumar A, Kumar A, Lata C, Kumar S (2016) Eco-physiological responses of Aeluropus lagopoides (grass halophyte) and Suaeda nudiflora (non-grass halophyte) under individual and interactive sodic and salt stress. South Afr J Bot 105:36–44
Kumar A, Kumar A, Kumar P et al (2018a) Effect of individual and interactive alkalinity and salinity on physiological, biochemical and nutritional traits of Marvel grass. Indian J Exp Biol 56:573–581
Kumar A, Kumar A, Lata C et al (2018b) Effect of salinity and alkalinity on response of halophytic grasses Sporobolus marginatus and urochondra setulosa. Ind J Agri Sci 88(8):150–157
Kumar A, Mann A, Kumar A et al (2018c) Potential and role of halophyte crops in saline environments. Engineering practices for management of soil salinity. Apple Academic Press, pp 379–416
Kumar A, Kumar A, Mann A et al (2019a) Phytoamelioration of the salt-affected soils through Halophytes. –. In: Hasanuzzaman M et al (eds) Ecophysiology, Abiotic stress responses and utilization of Halophytes. Springer, Singapore, pp 313–326
Kumar A, Mann A, Lata C et al (2019b) Salinity-induced physiological and molecular responses of Halophytes. Research Developments in saline agriculture. Springer, pp 331–356
Kumar R, Singh A, Bhardwaj AK et al (2022) Reclamation of salt-affected soils in India: progress, emerging challenges, and future strategies. Land Degrad Dev 33(13):2169–2180. https://doi.org/10.1002/ldr.4320
Lata C, Soni S, Kumar N et al (2019) Adaptive mechanism of stress tolerance in Urochondra (grass halophyte) using roots study. Indian J Agri Sci 89:1050–1053
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-∆∆CT method. Methods 25:402–408
Mangalassery S, Dayal D, Kumar A et al (2017) Pattern of salt accumulation and its impact on salinity tolerance in two halophyte grasses in extreme saline desert in India. Indian J Exp Biol 55(8):542–548
Mann A, Kumar N, Lata C et al (2019) Functional annotation of differentially expressed genes under salt stress in Dichanthium annulatum. Plant Physiol Rep 24:104–111. https://doi.org/10.1007/s40502-019-0434-88
Mann A, Kumar A, Saha M et al (2019a) Stress induced changes in osmoprotectants, ionic relations, antioxidants activities and protein profiling characterize Sporobolus marginatus Hochst. Ex A. Rich. Salt tolerance mechanism. Indian J Exp Biol 57:672–679
Mann A, Kumar N, Kumar A et al (2021a) De novo transcriptomic data of salt tolerant halophytes Dichnathium annulatum (Forssk.) Stapf and urochondra setulosa (trin.) CE hubb. Data in Brief 39:107536
Mann A, Kumar N, Kumar A et al (2021b) De novo transcriptomic profiling of differentially expressed genes in grass halophyte Urochondra setulosa under high salinity. Sci Rep 11:1–14
Mann A, Lata C, Kumar N, Kumar A, Kumar A, Sheoran P (2023) Halophytes as new model plant species for salt tolerance strategies. Front Plant Sci 14:1137211. https://doi.org/10.3389/fpls.2023.1137211
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17:10–12
Mittler R, Vanderauwera S, Gollery M, Van Breusegem F (2004) Reactive oxygen gene network of plants. Trends Plant Sci 9:490–498
Mittova V, Volokita M, Guy M, Tal M (2000) Activities of SOD and the ascorbate-glutathione cycle enzymes in subcellular compartments in leaves and roots of the cultivated tomato and its wild salt-tolerant relative Lycopersicon pennellii. Physiol Plant 110:42–51
Mohammadi Z, Rastegar S, Abdollahi F, Hosseini Y (2018) Morphological and antioxidant enzymatic activity responses of sapodilla rootstock to salinity stress. J Plant Process Funct 6:23–28
Nakano Y, Asada K (1981) Hydrogen peroxide is scavenged by ascorbate-specific peroxidase in spinach chloroplasts. Plant Cell Physiol 22:867–880
Panda A, Rangani J, Kumar Parida A (2019) Cross talk between ROS homeostasis and antioxidative machinery contributes to salt tolerance of the xero-halophyte Haloxylon salicornicum. Environ Exp Bot 166:103799. https://doi.org/10.1016/j.envexpbot.2019.103799
Pooja D, Nandwal AS, Chand M et al (2019) Varietal variation in physiological and biochemical attributes of sugarcane varieties under difference soil moisture regimes. Indian J Exp Biol 52:721–732
Prasad G, Mittal S, Kumar A, Chauhan D, Sahu TK, Kumar S, Singh R, Yadav MC, Singh AK (2022) Transcriptome analysis of bread wheat genotype KRL 3–4 provides a new insight into regulatory mechanisms associated with sodicity (high pH) tolerance. Front Genet. 2022; 12:782366. https://doi.org/10.3389/fgene.2021.782366
Qi YC, Liu WQ, Qiu LY et al (2010) Overexpression of glutathione S-transferase gene increases salt tolerance of Arabidopsis. Russian J Plant Physiol 57:233–240
Raja V, Wani UM, Wani ZA et al (2021) Pyramiding ascorbate–glutathione pathway in Lycopersicum esculentum confers tolerance to drought and salinity stress. Plant Cell Rep 1–19
Rao MV, Watkins CB, Brown SK, Weeden NF (1998) Active oxygen species metabolism in ‘White Angel’ x ‘Rome Beauty’ apple selections resistant and susceptible to superficial scald. J Am Soc Hort Sci 123:299–304
Sahu BB, Shaw BP (2009) Isolation, identification and expression analysis of salt-induced genes in Suaeda maritima, a natural halophyte, using PCR-based suppression subtractive hybridization. BMC Plant Biol 9:69
Salbitani G, Bottone C, Carfagna S (2017) Determination of reduced and total glutathione content in extremophilic microalga Galdieria phlegrea. Bio-protocol 7:e2372–e2372
Sharma V, Kumar N, Verma A, Gupta VK (2013) Exogenous application of brassinosteroids ameliorates salt-induced stress in mung bean seedlings. LS: Int J Life Sci 2:7–13
Sheoran P, Sharma R, Kumar A, Singh RK, Barman A, Prajapat K, Kumar S, Sharma PC (2022) Climate resilient integrated soil-crop management (CRISCM) for salt affected wheat agri-food production systems. Sci Total Environ. Sep 1;837:155843. https://doi.org/10.1016/j.scitotenv.2022.155843
Sinha AK (1972) Colorimetric assay of catalase. Analy Biochem 47:389–394. https://doi.org/10.1016/0003-2697(72)90132-7
Souid A, Bellani L, Magné C et al (2018) Physiological and antioxidant responses of the sabkha biotope halophyte Limonium delicatulum to seasonal changes in environmental conditions. Plant Physiol Biochem 123:180–191
Tsukagoshi H, Suzuki T, Nishikawa K et al (2015) RNA-seq analysis of the response of the halophyte, Mesembryanthemum crystallinum (ice plant) to high salinity. PLoS ONE 10:e0118339
Wang J, Li B, Meng Y et al (2015) Transcriptomic profiling of the salt-stress response in the halophyte Halogeton glomeratus. BMC Genomics 16:1–14
Yamamoto N, Takano T, Tanaka K et al (2015) Comprehensive analysis of transcriptome response to salinity stress in the halophytic turf grass Sporobolus virginicus. Front Plant Sci 6:241
Yi Z, Li S, Liang Y et al (2018) Effects of exogenous spermidine and elevated CO2 on physiological and biochemical changes in tomato plants under iso-osmotic salt stress. J Plant Growth Reg 37:1222–1234
Zang W, Miao R, Zhang Y et al (2021) Metabolic and molecular basis for the salt and alkali responses of Suaeda corniculata. Environ Exp Bot 192:104643
Zhang H, Zhai N, Ma X et al (2021) Overexpression of OsRLCK241 confers enhanced salt and drought tolerance in transgenic rice (Oryza sativa L). Gene 768:145278
Acknowledgements
We gratefully acknowledge the Indian Council of Agricultural Research (ICAR) for providing us the financial support to conduct the work. We are also thankful to the Director, ICAR-Central Soil Salinity Research, Karnal, Haryana, India for imparting required facilities for carrying out the research work.
Funding
This project was supported by the Indian Council of Agricultural Research (ICAR) - National Agricultural Science Fund (NASF), New Delhi, India (Grant no ABP-6027).
Author information
Authors and Affiliations
Contributions
A.M. formulated the experiments, organized and prepared the main manuscript, supervision of project. N.K. performed QC, gene validation and bioinformatic analysis, wrote the original manuscript draft. C. L. performed total RNA isolation and physiological enzyme analysis. Ar. K. discussed data presentation design. B.L.M. helped in maintaining salinity throughout the experiment and A.K. interpreted results and performed SAS.
Corresponding author
Ethics declarations
Ethical approval and consent to participate
This article does not involve any studies on humans/animals conducted by any of the authors.
Consent for publication
All authors have read and approved the final manuscript.
Conflict of interest
All the authors declare no competing interests.
Additional information
Communicated by Zhong-Hua Chen.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10725_2023_1085_MOESM1_ESM.jpeg
Supplementary Material 1: Supplementary Fig. 1. Important pathways operative in ROS system in U. setulosa and D. annulatum at salinity levels of EC 30, 40 & 50 dSm− 1
10725_2023_1085_MOESM2_ESM.jpeg
Supplementary Material 2: Supplementary Fig. 2. GO enrichment of important terms in both the plants at salinity levels of EC 30, 40 & 50 dSm− 1
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mann, A., Kumar, N., Lata, C. et al. Physiological and differential gene expression reveals a trade-off between antioxidant capacity and salt tolerance in Urochondra setulosa and Dichanthium annulatum. Plant Growth Regul 102, 555–570 (2024). https://doi.org/10.1007/s10725-023-01085-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-023-01085-y